Summary: A brand new technique dubbed STARmap PLUS was utilized to trace and map modifications in tau and amyloid beta within the mind as Alzheimer’s illness progressed in mouse fashions.
Source: MIT
A standard signal of Alzheimer’s illness is the extreme buildup of two forms of protein within the mind: tangles of tau proteins that accumulate inside cells, and amyloid-β proteins that type plaques exterior the cells. Researchers don’t know the way these protein deposits are associated to the opposite main hallmark of the illness: the demise of neurons within the mind.
A research by scientists on the Broad Institute of MIT and Harvard printed at present in Nature Neuroscience hints at some solutions to this query.
The group used a brand new technique they developed to disclose how mind cells situated close to these proteins change because the illness progresses in a mouse mannequin of Alzheimer’s. The approach, referred to as STARmap PLUS, is the primary to concurrently map gene expression of particular person cells and their location, in addition to the spatial distribution of particular proteins in intact tissue samples.
The researchers used their strategy to review mind tissue from the Alzheimer’s mouse mannequin at two completely different levels of the illness and at excessive spatial decision. In the sooner stage, they noticed a central core of amyloid plaque surrounded by a sort of immune cell within the mind referred to as microglia, that are identified to play a job in Alzheimer’s. The microglia that have been nearer to the plaques confirmed genetic signatures which have been linked to neurodegeneration.
The scientists additionally discovered outer shells of two different forms of mind cells that emerged later within the illness. This core-shell construction and variations in gene expression of cells surrounding the proteins give scientists a clearer image of how cells reply to the protein deposits within the mind — insights that would someday assist scientists consider present Alzheimer’s remedies and develop new ones.
“From these kinds of studies, you can infer what’s going on in a far more detailed way than you could if you just looked at cells from dispersed tissue samples, that don’t have their spatial context anymore,” mentioned Morgan Sheng, co-senior creator of the research, a core institute member and co-director Stanley Center for Psychiatric Research on the Broad, and a professor of neuroscience at MIT. “This is a new dimension of transcriptomics, and I think it’s going to be really impactful.”
The research builds on a earlier model of the approach referred to as STARmap, developed by Xiao Wang, who’s a co-senior creator on the research, a core institute member and Merkin Fellow on the Broad, and a professor of chemistry at MIT.
“This is an exciting improvement on STARmap because we can now co-map the entire transcriptome together with proteins in the same tissue slices, and many diseases involve changes in protein localization and post-transcriptional modifications,” mentioned Wang.
The venture can also be a collaboration with Genentech scientists and was led by co-first authors from the Stanley Center: Hu Zeng, a postdoctoral fellow; Jiahao Huang, a graduate scholar; and Haowen Zhou, a visiting researcher.
Making a map
To analyze tissue samples utilizing STARmap PLUS, Wang’s group used molecular probes to detect particular mRNAs and amplify them as DNA sequences. They additionally used antibodies to label and establish particular proteins. They then chemically handled the tissue to anchor the DNA and proteins of their native positions inside a gel. Finally, they used in situ sequencing and imaging to create a three-dimensional map of the tagged proteins in addition to the expression of greater than 2,700 genes.
The scientists discovered that processes such because the mind’s inflammatory response and the differentiation of glial cells such because the microglia have been related to illness development. Though different researchers had beforehand noticed a core-shell construction round plaque, the brand new gene expression information revealed that the microglia have been extra “activated” to set off an inflammatory response nearer to the plaque.
The scientists say that this means that the microglia possible activate close to plaques, probably recruiting different cells to type the outer shells surrounding the plaques, slightly than activating far-off after which shifting nearer. Understanding when, the place, and the way microglia activate could possibly be an essential a part of deciphering their function within the illness.
Wang says {that a} key benefit of STARmap PLUS is that it collects each protein and gene expression data from a single pattern, making it simpler to align and examine completely different varieties of information at excessive decision. It may also detect options smaller than cells, which helps distinguish particular person cells even when they’re densely crowded collectively within the mind. STARmap PLUS can also be scalable and could possibly be tailored to map different proteins and even your complete transcriptome.
Beyond Alzheimer’s
The researchers say {that a} essential subsequent step might be to make use of the strategy to review Alzheimer’s development in human mind tissue samples. This will assist decide the extent to which mobile modifications that happen in mouse fashions characterize processes in Alzheimer’s sufferers.
In animal fashions, scientists may additionally use the strategy to reply questions on new therapy methods. For instance, if antibodies are capable of attain and clear plaques, do the close by microglia return to their unactivated states and transfer away from the plaques? Does eliminating plaque or inactivating microglia forestall close by neurodegeneration?
STARmap PLUS may additionally assist researchers perceive different illnesses, equivalent to most cancers, to study extra, for instance, about how immune cells assault tumors. The technique may additionally contribute to research on schizophrenia and different mind problems.
“There are mouse models of psychiatry, where we know from other studies that there are many different things happening in different parts of the brain,” Sheng mentioned. “It’ll be just gorgeous to be able to see it all in one swoop.”
Funding: This work was supported partly by the Searle Scholars Foundation, the Stanley Center for Psychiatric Research, and the Merkin Institute.
About this Alzheimer’s illness and mind mapping analysis information
Author: David Cameron
Source: MIT
Contact: David Cameron – MIT
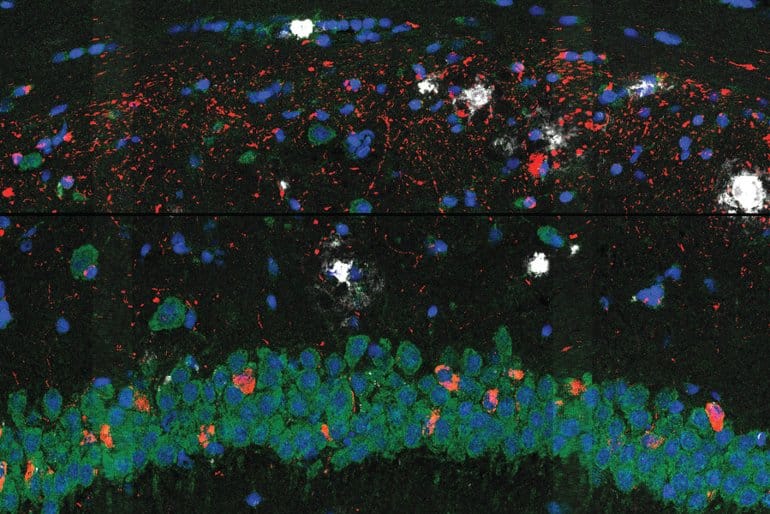
Image: The picture is credited to Zeng H, Huang J, Zhou H, et al
Original Research: Closed entry.
“Integrative in situ mapping of single-cell transcriptional states and tissue histopathology in a mouse model of Alzheimer’s disease” by Zeng H, Huang J, Zhou H, et al. Nature Neuroscience
Abstract
Integrative in situ mapping of single-cell transcriptional states and tissue histopathology in a mouse mannequin of Alzheimer’s illness
Complex illnesses are characterised by spatiotemporal mobile and molecular modifications which may be tough to comprehensively seize.
However, understanding the spatiotemporal dynamics underlying pathology can make clear illness mechanisms and development.
Here we introduce STARmap PLUS, a technique that mixes high-resolution spatial transcriptomics with protein detection in the identical tissue part.
As proof of precept, we analyze mind tissues of a mouse mannequin of Alzheimer’s illness at 8 and 13 months of age.
Our strategy gives a complete mobile map of illness development. It reveals a core–shell construction the place disease-associated microglia (DAM) carefully contact amyloid-β plaques, whereas disease-associated astrocyte-like (DAA-like) cells and oligodendrocyte precursor cells (OPCs) are enriched within the outer shells surrounding the plaque-DAM complicated.
Hyperphosphorylated tau emerges primarily in excitatory neurons within the CA1 area and correlates with the native enrichment of oligodendrocyte subtypes.
The STARmap PLUS technique bridges single-cell gene expression profiles with tissue histopathology at subcellular decision, offering a instrument to pinpoint the molecular and mobile modifications underlying pathology.






Discussion about this post